Molecular diagnostics is coming of age in parasitology and may help in diagnosing and managing disease. This includes determining which isolates of coccidia should be used in immunization with live vaccines. It may also be used to track the sources of outbreaks of blackhead and identify responsible vectors.
Polymerase Chain Reaction
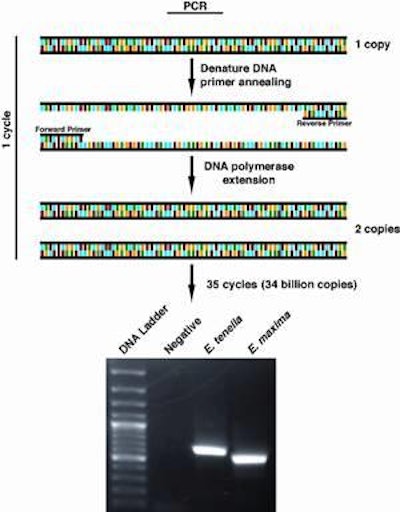
This can be accomplished through the use of PCR, the acronym for Polymerase Chain Reaction, which has virtually become a household word as people watch TV crime programs. Most people don’t realize that this same tool is being applied to diagnose human diseases ranging from HIV to Salmonella to amebic dysentery.
The advantages of PCR diagnostics has not been lost on the poultry industry, with most viral diseases now being detected using the technology and tests for other pathogens under development. The success of PCR has created a revolution in diagnostics and research labs. PCR promises to play a special role in poultry parasitology.
PCR’s role in parasitology
How does PCR technology apply to coccidiosis and other parasitic diseases?
Some coccidia are easily recognized by examination of infected animals and microscopic examination of oocysts, the infective reproductive cells. This works well for species like Eimeria tenella (bloody cecal coccidiosis), E. acervulina (upper intestinal coccidiosis) or E. necatrix (bloody intestinal coccidiosis).
People often wonder why E. hagani, E. mitis, E. praecox or E. mivati are not often diagnosed and assume that they must be rare. The fact is, the only way to specifically identify these species is a long ordeal in the laboratory involving infection of live birds, measuring the time before shedding of oocysts in the droppings, measurement of hundreds of oocysts, looking at lesions, and conducting cross/immunization experiments. Usually, the information that would be obtained is no longer useful.
Diagnosing species
The problem is even worse with turkey coccidia, pheasants and other gamebirds, where so few studies have been done in recent years that most workers have little experience in diagnosing species. Regulatory authorities often opine that some of the species do not exist or cause disease, simply because no one has diagnosed them recently. The rapidity and reliability of the PCR test for these parasites will be welcomed in diagnostic labs.
Replacement for trained parasitologists?
The number of scientists and veterinarians working with coccidia has shrunk considerably in recent years. Without a pool of experienced workers to recognize the different species, there is no way to conduct meaningful research on these organisms and determine which of the species are responsible for serious outbreaks of disease. This also hinders how we determine which isolates of coccidia should be used in drug tests or immunization with live vaccines. Strains vary in virulence, immune types, and other characters. This is where PCR can fill the void.
Primers allow PCR identification
Because of sequence variation between each species of coccidia, primers can be designed that allow for PCR identification. For much of this work, the ITS1 (for internal transcribed spacer) and ITS2 region found between structural ribosomal RNAs (rRNA) are used (Figure 2A). This nonfunctional region in the genome has limited pressure to preserve its sequence integrity allowing changes to occur between closely related species and even strains within species.
Studies monitor field species
As seen in Figure 2B, the ITS1 regions for E. acervulina, E. maxima and E. tenella are somewhat variable. This type of divergence allows PCR primers to be constructed that are specific to each coccidia species.
Recently, PCR was applied to the question of speciation of turkey coccidia, with the result that all the known species were found. Similar studies have been used to monitor the progression of chicken coccidiosis in the field in the USA, Holland, India, Australia, China and other countries.
Bar code for detection of virulence in parasites?
Considerable variation is known between strains of parasites that are within the same species. Taking advantage of the widely variable ITS regions, multiple strains of E. maxima have been identified (Figure 2C). These strains differ in virulence and immunogenicity. In this manner, the ITS sequence acts as a bar code that can be scanned by PCR to type these organisms.
Another example is the study of blackhead organisms (Histomonas) for differences associated with virulence, host and source. Considerable difference in virulence is known, particularly in chickens.
Identifying and tracking blackhead
In a recent test, we found differences in the ITS1 and ITS2 sequences between isolates acquired from Georgia, North Carolina, Arkansas and California, and particularly between chickens and turkeys. Further studies are under way to correlate these differences with virulence and to use molecular techniques to identify the genes responsible for this trait.
There is considerable interest in ‘tracking’ the sources of outbreaks of blackhead and identification of responsible vectors. Of no small consequence, these studies also identified possible cases of misdiagnosis. Blackhead is sometimes confused with other protozoal or bacterial infections because of similarity in gross lesions. PCR is valuable in all of these applications.
More applications for the future?
These are but a few examples of how PCR can be applied in study of coccidiosis and blackhead in diagnostic and research labs. As in many other fields, determined investigators are at work in laboratories seeking new facets of this latest tool in the fight against disease.