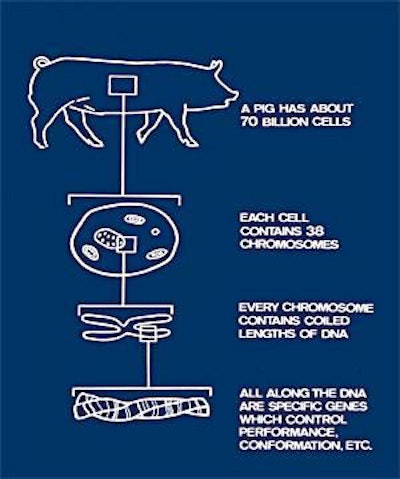
As well as being the most important meat animal in the world, the pig is used as a model for human surgery, health studies and potential organ transplantation. As a result, the publication of the first draft of the pig genome in November 2009 heralded much interest in the world’s press. Uniquely, this was the only genome-sequencing project in the world that was both a public-private partnership and a global collaborative effort, and the total cost was estimated to have been some US$24.3 million.
The funding and technical support for the project came from the EU, France, Japan, Korea, the Netherlands, the UK and US and involved co-ordination and co-operation between several international teams of scientists and genome-sequencing centers. A key requirement of participation was that the findings be made public, with no proprietary interests allowed.
Multiple benefits
The draft sequence, which is about 98% complete, allows researchers to pinpoint genes that are useful to pork production or are involved in immunity or other important physiological processes. It has the potential to increase genetic progress as well as offering an insight into pig disease and immunity traits and will assist in efforts to preserve the global heritage of rare, endangered and wild pigs.
It also will be important for the study of human health because pigs are very similar to humans in their physiology, behaviour and nutritional needs.
Traits of economic importance
To date, researchers have identified several genes or DNA regions that are associated with traits of economic importance including reproduction, growth, lean and fat quantity, meat quality traits and disease resistance.
A number of gene and marker tests are now available commercially from genotyping service companies. Examples are CAST (meat quality), ESR and EPOR (litter size), FUT1 (E. coli disease resistance), HAL (halothane – meat quality, stress), IGF2 (carcase), MC4R (growth and fat), PRKAG3 (meat quality), RN (meat quality).
Many commercial breeding companies are now using this information combined with traditional performance information to improve pig production using a technique called marker assisted selection (MAS). However, the full gene sequence is still not complete and further work is required to allow significant and real commercial benefits to continue to flow.
Increasingly we are learning that gene interactions (and their interaction with the environment) are very complex. The simple view of 10 years ago that the publication of the porcine genome would unlock numerous advances is partly untrue. A very simple analogy would be that 10 years ago we had a Ferrari, but if there was no written manual then driving it to its potential and fixing its faults would be difficult. Now we have a manual (with several pages missing) so we are better off than before, but still need more information and experience to get the full potential from the machine.
In addition to the recent developments in pig genome sequencing there has also been the release of single-nucleotide polymorphism, or SNP, chips. These chips are designed to genotype individual pigs for some 60,000 SNPs at any one time.
Current costs are too high for use by all except the largest breeding companies, but these costs are reducing fast so that the tool will be available to a wide range of breeders before too long. It turn, this will allow the use of genomic selection, which is the prediction of a selection candidate’s breeding value based on the SNP data.
These predictions, together with the completion of the pig genome sequence, offer the possibility to select pigs for traits that, to date, have been difficult to improve, such as robustness and general disease resistance, as they look at the relationships between animals at the DNA level, rather than prediction on the average performance from test and pedigree data.